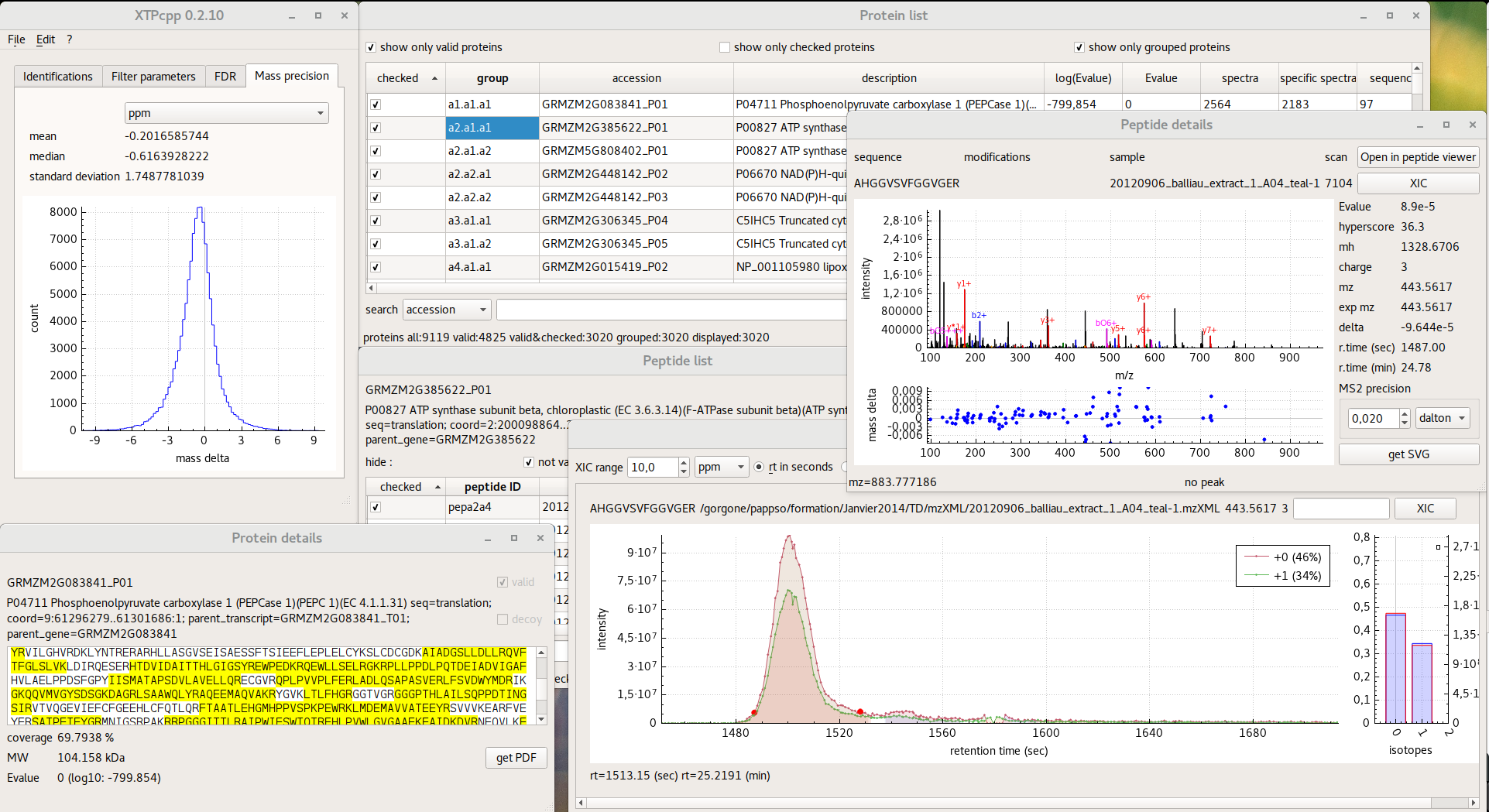
Here is the changelog since last announced 0.2.32 version :
- Small screen adaptation. A lift is available if the screen is too small to be able to click on OK button.
- New choice in the computation of XIC intensities. “sum” computes the sum of intensities inside the precision range. “max” will take only the maximum of intensitie.
- New screen to edit global settings (edit => settings). 2 methods to extract XICs are available. “buffered” is very fast but requires some extra space on the hard disk. “direct” is a bit slower but does not require extra space.
- Better ODS exports. new column to display mass delta in “ppm” in the “spectra” sheet.
- Modifications in “protein list” and “peptide list” windows. A new menu is available to select “valid”, “checked” and “grouped” proteins/peptides. A direct export in tabulated sheet of the view is available. Flanking Nter and Cter regions are now displayed.
- Better X!Tandem import filter to take into account “point mutation”.
- New filter “peprepro” to select valid peptides. It allows to consider peptides only if they are seen in one or more MS runs.
- Bette PROTICdbML support
- New controls to help the use of X!Tandem
Many thanks to Solenne Chardonnet, Cédric Pionneau, Valérie Briard-Bion, Julien Jardin and the PAPPSO team for their contributions.